We’ll be back soon!
Sorry for the inconvenience but we’re performing maintenance at 5:00 a.m. on Monday, September 18, 2023 through 6:00 p.m. on Wednesday, September 20, 2023 (PST). We’ll be back online shortly!
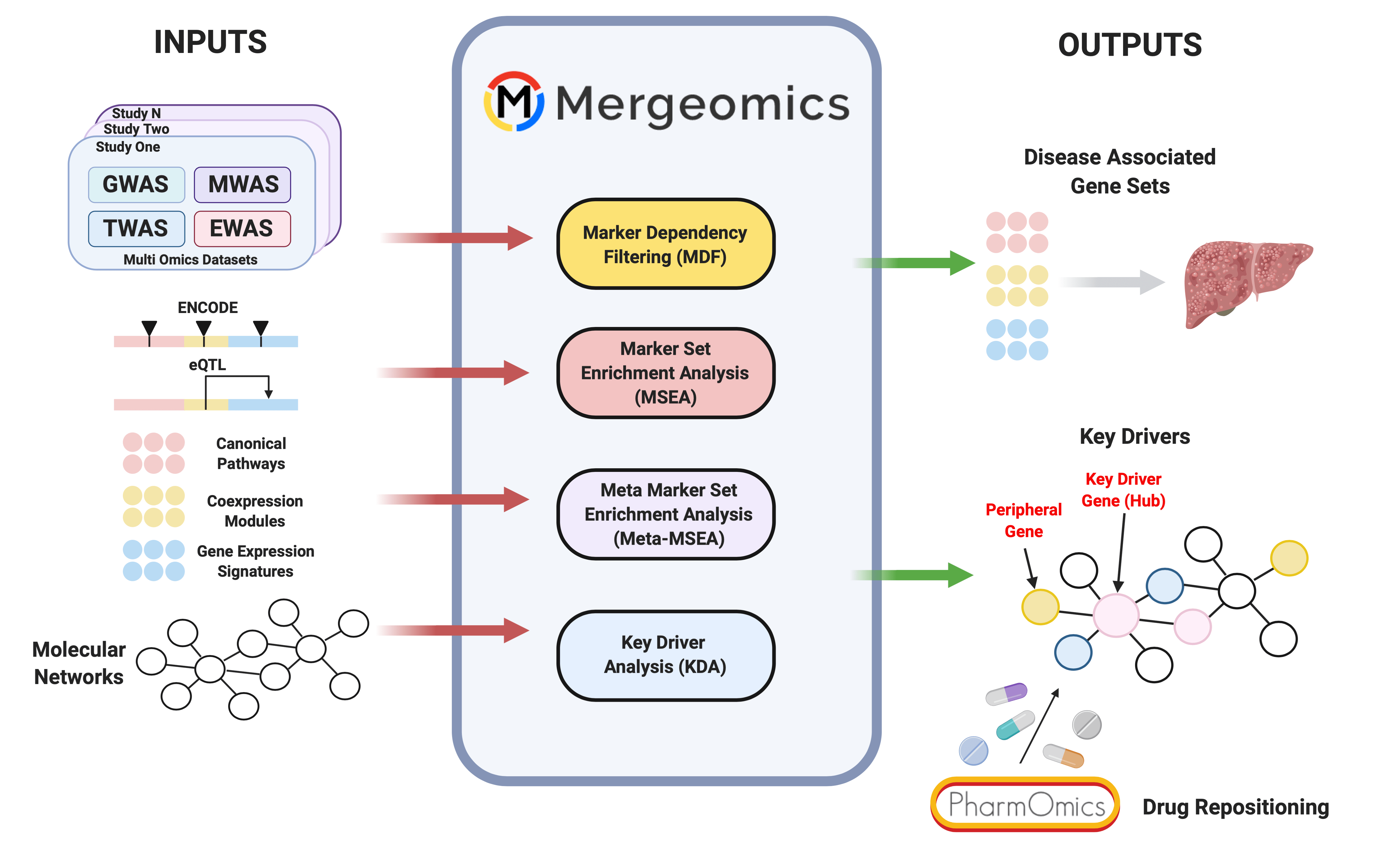
— Mergeomics & Pharmomics Team
Don't show this again!